tyrosinemia type 1
Tyrosinemia Type 1 is an autosomal recessive disorder that is characterized by the inability to break down the amino acid tyrosine within the body. Patients with Tyrosinemia Type 1 have a mutation within the FAH gene (located on chromosome 15) that encodes for the enzyme fumarylacetoacetate hydrolase. Without this functioning enzyme, the final step in the tyrosine degradation process cannot be complete, and succinylacetone builds up in the body, leading many health problems throughout the body, mainly within the liver and kidneys. The main symptom of this disorder is a progression of a healthy liver to one with cirrhosis and eventually liver cancer. Other symptoms include rickets, jaundice, and the inability to gain weight.
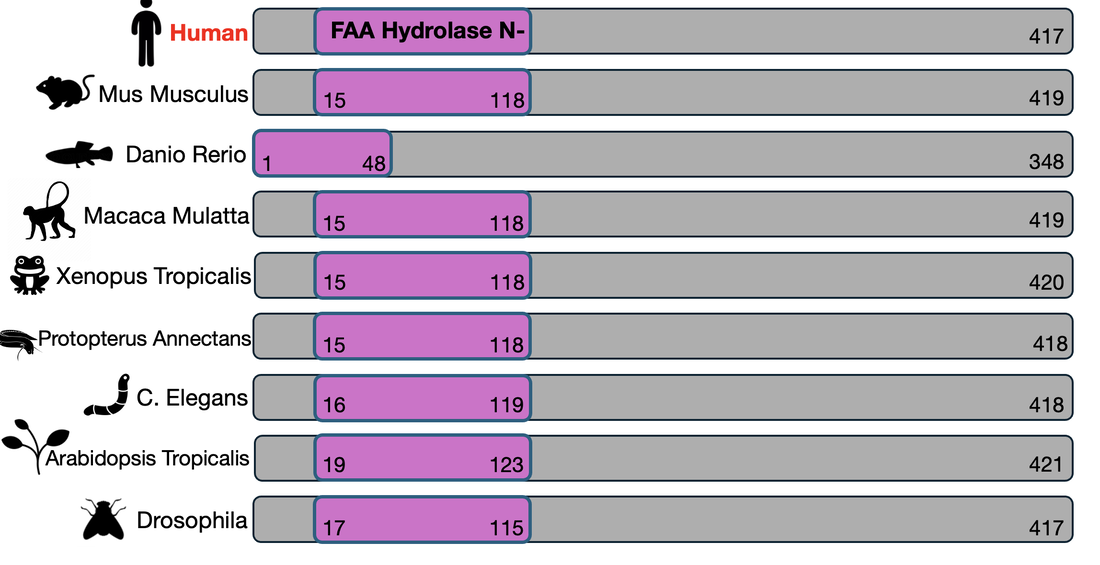
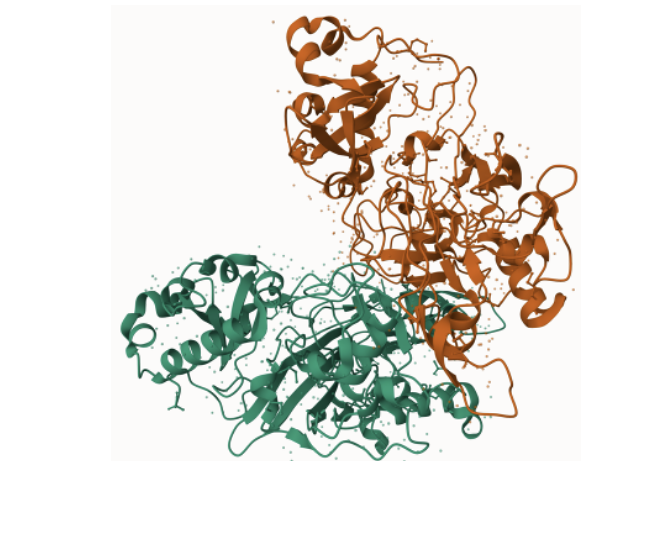
fumarylacetoacetate hydrolase (faa) domain
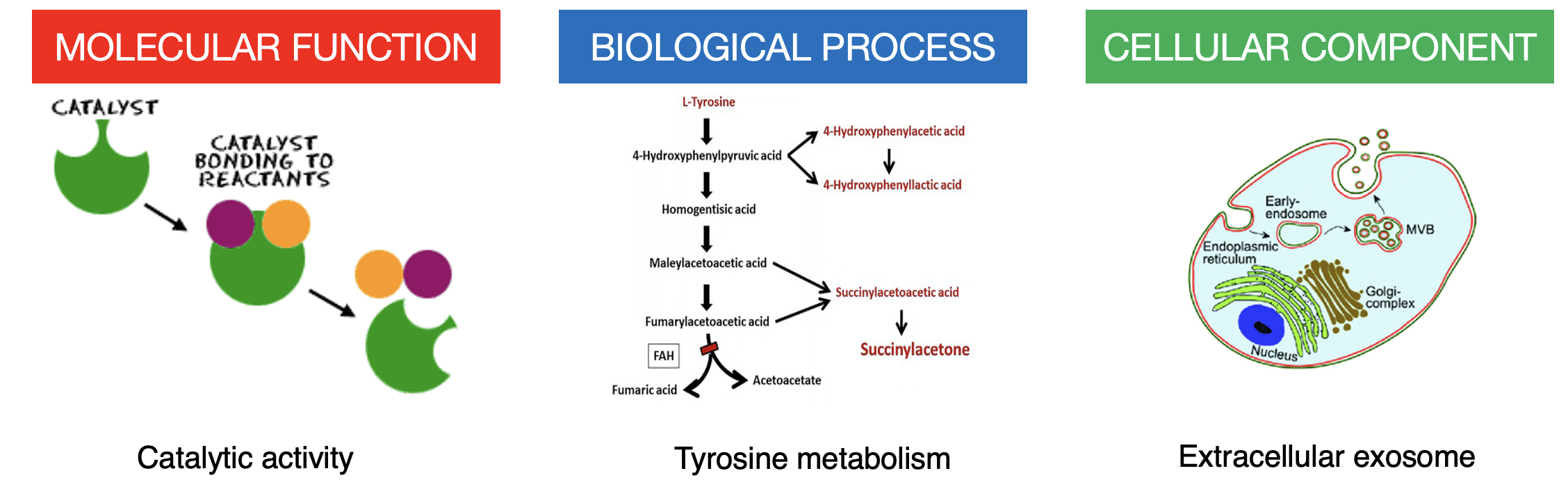
gene ontology
Gene ontology works to classify genes by their actions within a cell. The molecular function of FAA is catalytic activity, meaning it serves as a catalyst, speeding up a reaction within the cell. The biological process is the tyrosine metabolism pathway, serving as the last step in the pathway. The cellular component is function within extracellular exosomes. Placing these togther, it is seen that the FAH gene encodes for an enzyme that acts as a catalyst for the last step in the tyrosine metabolism pathway within the extracellular exosome.
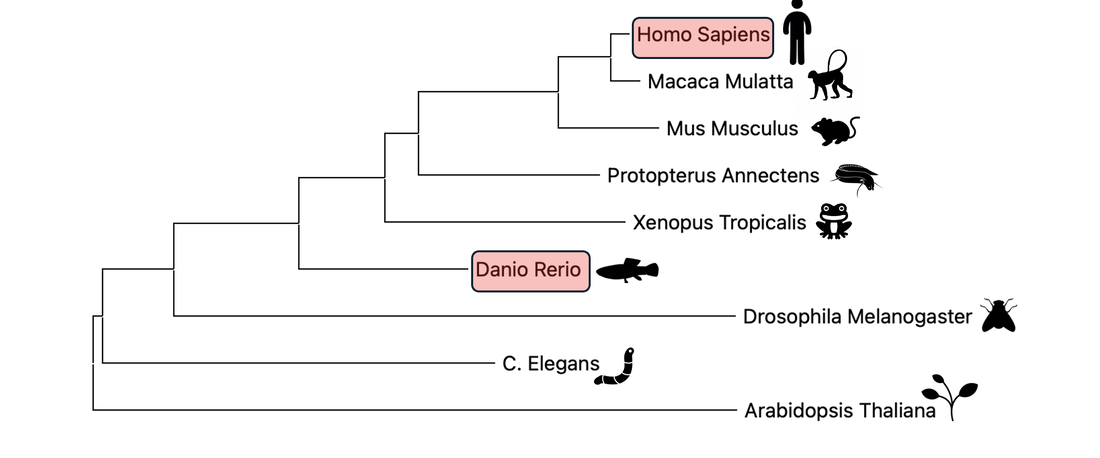
conservation and phylogeny
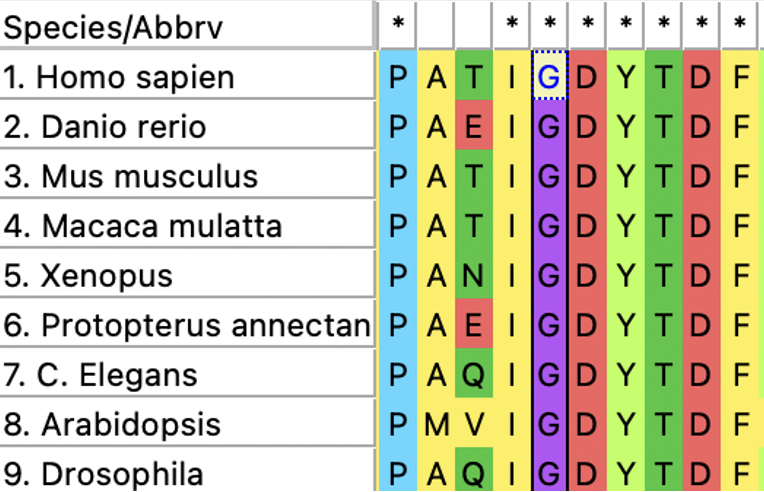
The FAA domain is highly conserved across many species. It's conservation can be seen in mammals, amphibians, nematodes, and even plants, highlighting it's evolutionary importance. These relationships are further visualized with a phylogenetic tree.
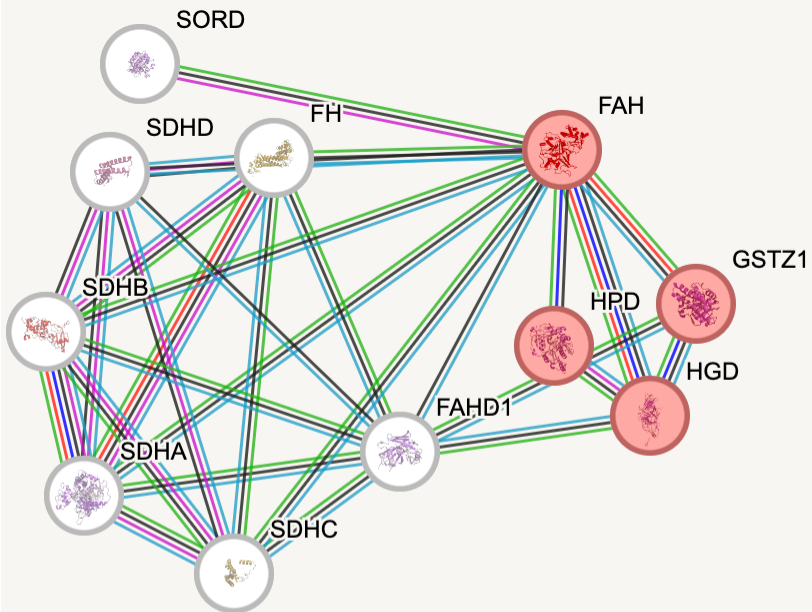
faa protein interactions
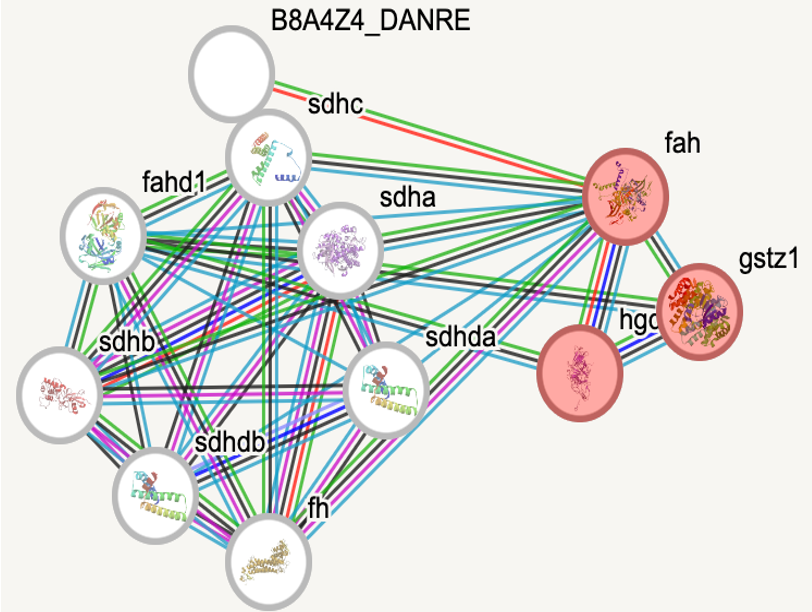
Protein interaction networks are an important tool for understanding the difference between a cell that is healthy and functioning and one that is not, as these protein-protein interactions are essential to nearly every process. Looking at these protein interaction networks for both homo sapiens and danio rerio, which will be used as the model organism for this research, it is seen that the protein interactions involved in each organism are very similar. The only key difference to note when looking at the proteins involved in tyrosine catabolism (in red) is that there is a protein that humans have that danio rerio do not.
danio rerio will function as the model organism
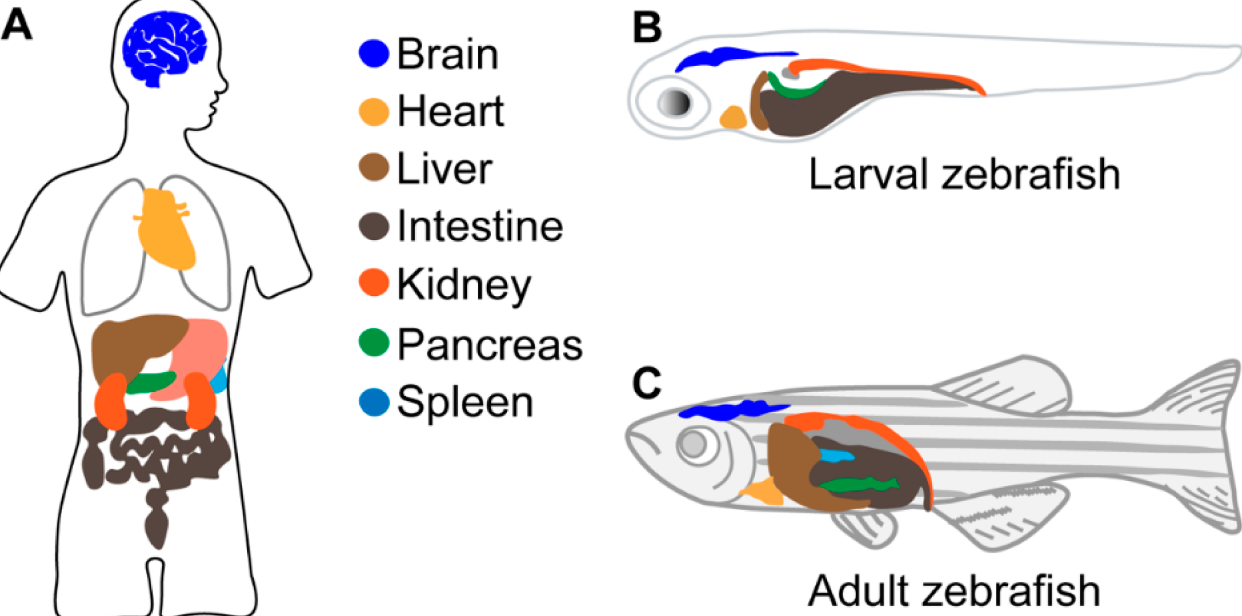
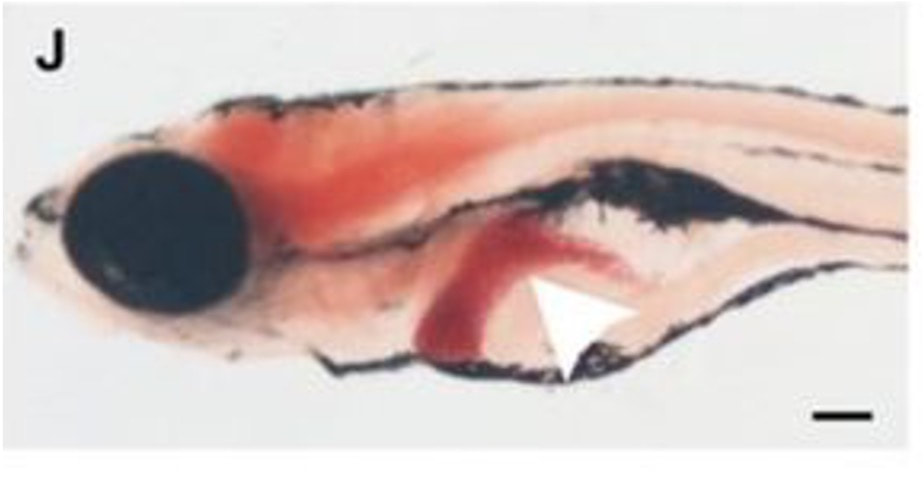
Zebrafish act as an excellent model organism for modeling different liver diseases in humans. With the previously observed conservation of the FAA domain and the similarities between organ systems within humans and zebrafish, they serve as a perfect model organism for this research. The transparency also allows for the liver to be seen externally and studied, which becomes very useful in research such as this one.
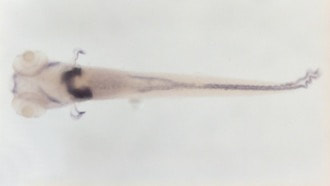
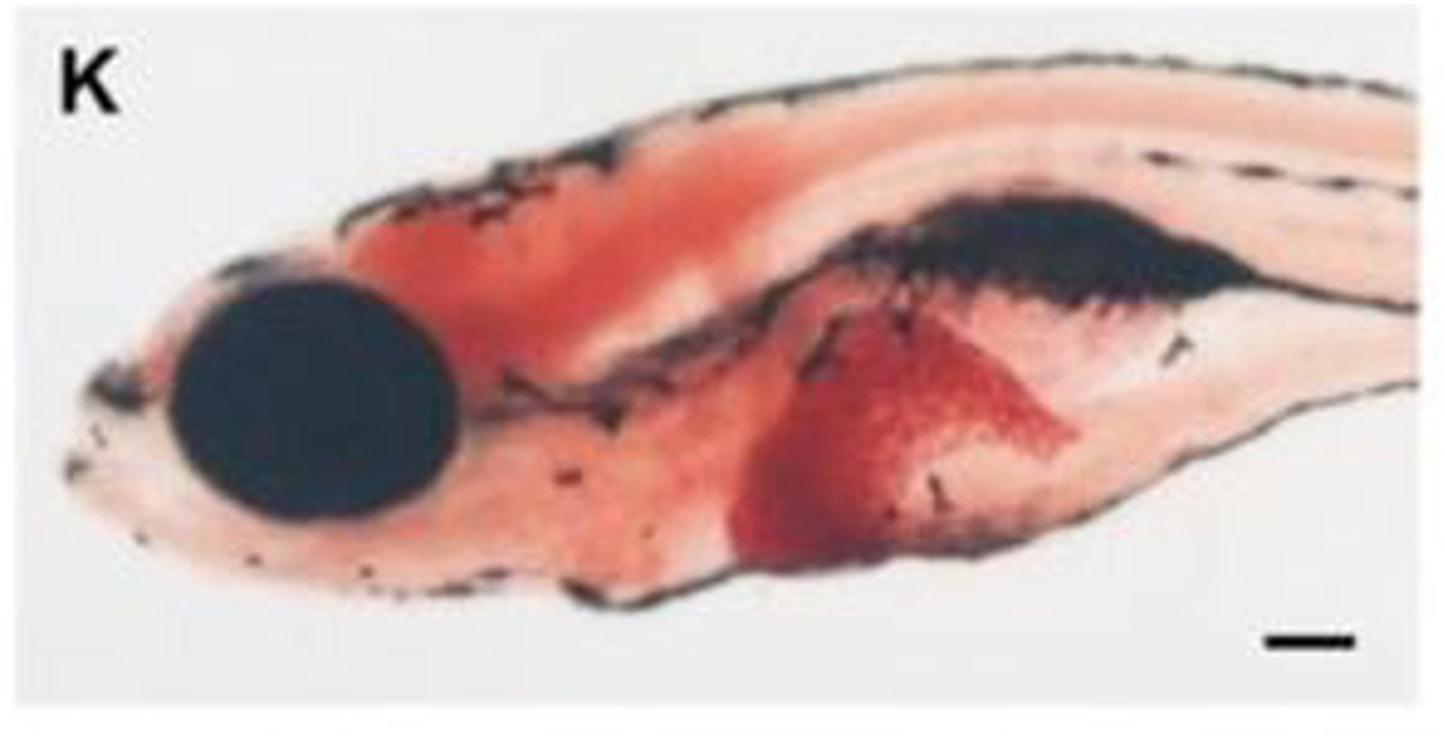
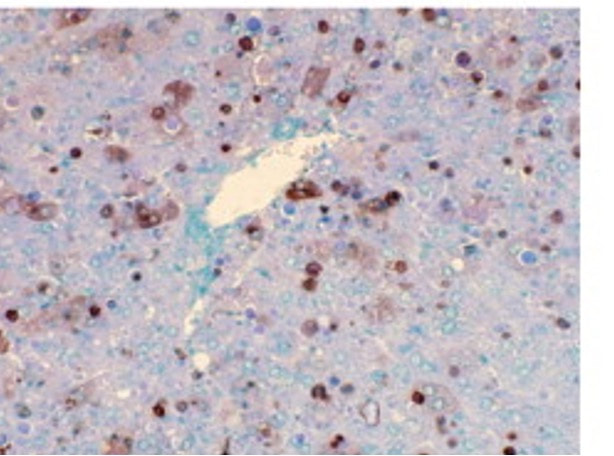
When a normal zebrafish is taken, the FAH gene is functioning and a normal-sized, healthy liver is observed in the fish. Looking at the apoptotic assay of this fish, there is some apoptosis, which is normal as some cells in the body do not function properly, however there is overall not a lot of apoptosis seen. When a zebrafish with a mutation in the FAH gene is observed, the liver can be seen to be much smaller and less healthy due to the extreme increase in apoptosis, which can be seen on the apoptotic assay for this zebrafish. This relationship is shown below and is a great reason why danio rerio serve as the best model organism for this research.
gap in knowledge
|
While Tyrosinemia Type 1 is a well studied disorder, there is still a lot unknown about the effects this has on the function of the body. Specifically, it remains unknown how the lack of a functioning FAA enzyme leads to the increase in hepatocyte apoptosis that is observed in patients exhibiting liver problems with this disorder.
|
I hypothesize that the FAH gene regulates a key process involved in healthy cell progression and without this functioning gene, hepatocyte cells are stalled in the cell cycle, leading to increased apoptosis and in turn, cirrhosis of the liver.
primary research goal
The primary goal of this research is to further explore how mutations in FAA lead to increased apoptosis of cells in the liver. In order to do this, three aims were crafted that utilize genomic and proteomic approaches to investigate this overall goal.
Aim 1 : Identify conserved amino acids of faa necessary for healthy cell progression.
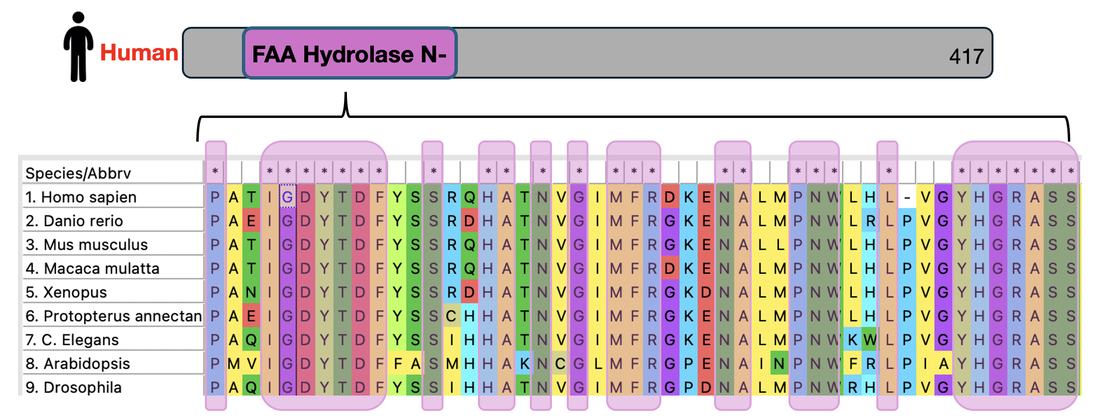
First, Pfam will be used to identify protein domains on the FAH gene. Using this domain and using Clustal Omega to create an amino acid sequence alignment, conserved amino acids across common homologous model organisms are found and highlighted below.
Choosing three of these conserved amino acids at random, amino acids S23, G31, and L48 were chosen to be mutated for this experiment.
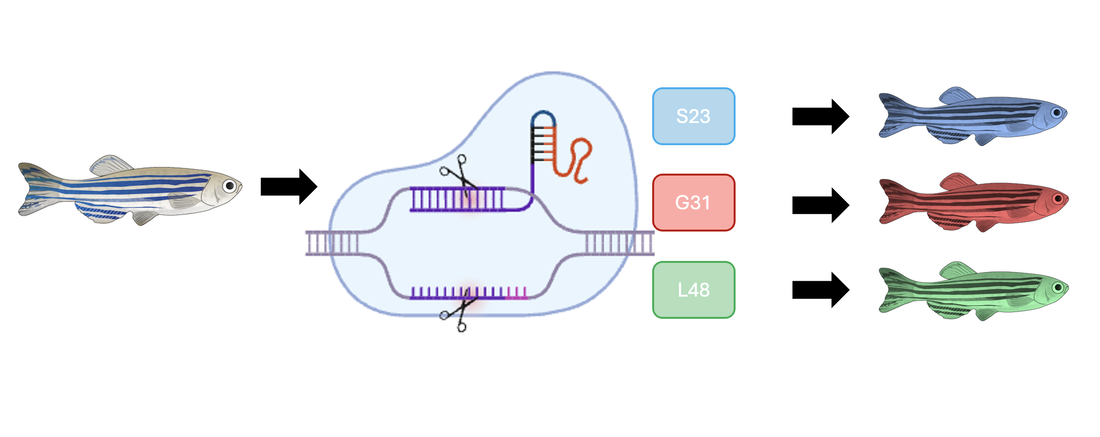
Using CRISPR, these amino acids will be mutated and create 3 lines of differently mutated fish, the blue fish representing the S23 mutation, the red fish representing G31 mutation, and the green fish representing the L48 mutation.
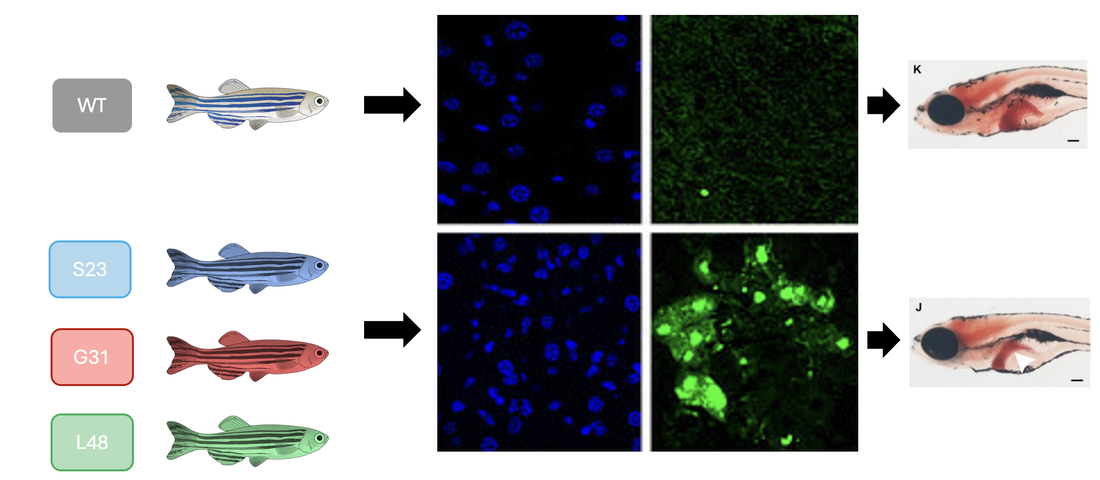
Using a TUNEL assay to assess apoptosis, the WT control group will be compared to each mutant fish through screening as well as phenotypic analysis. Below, the WT control group is shown to have low levels of apoptosis and a healthy, normal-sized liver while the fish with the mutated amino acids have an extreme increase in apoptosis as well as a decrease in liver size and health.
Overall, this research would reveal amino acids that are important for healthy cell progression in liver cells and will be useful in analyzing future research regarding how to stop these mutations as well as possible gene editing options that could fix these mutations.
Aim 2 : identify differentially expressed genes in wt and mutant fah hepatocyte cells.
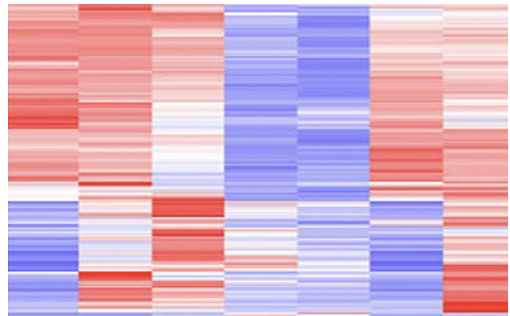
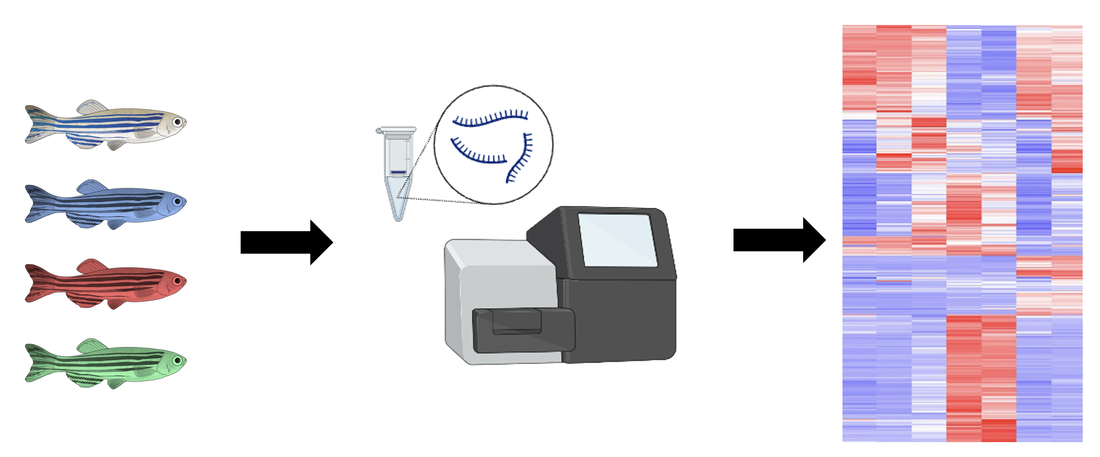
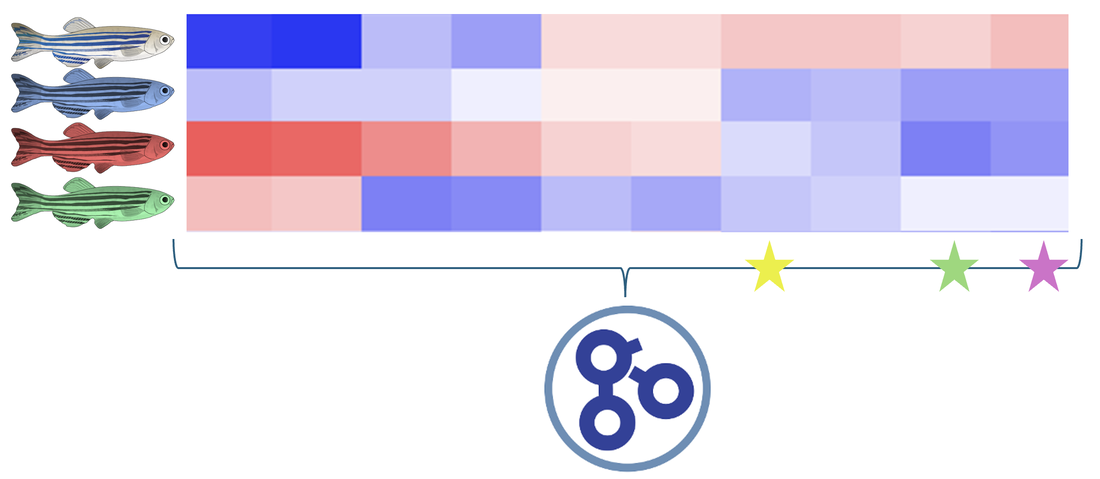
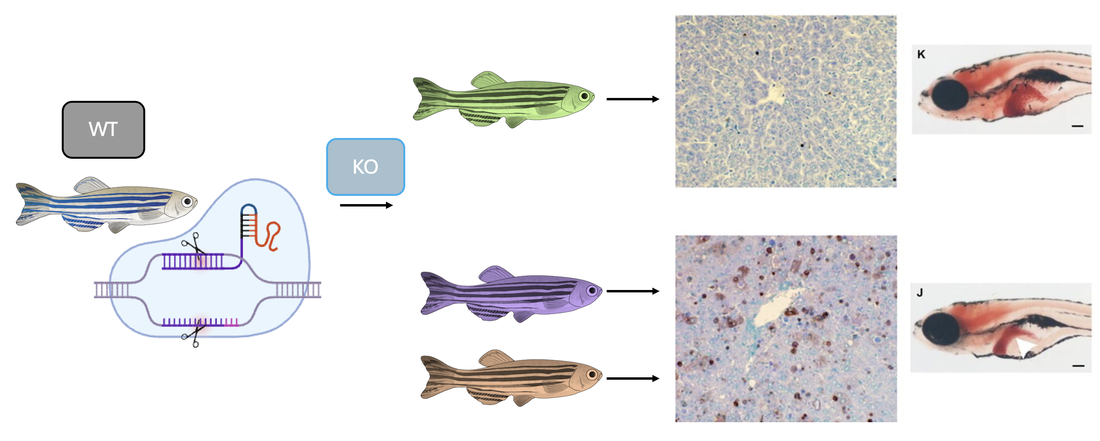
The mutant zebrafish from the previous aim will be used for further research regarding gene expression. RNA will be extracted from each mutant fish as well as a WT control group and ran through RNA-sequencing to produce a heat map showing gene expression.
Sorting this heat map for gene ontology (GO) term tyrosine catabolism pathway, 3 genes were selected for further evaluation. These genes were selected because they were downregulated in the WT control group, and found upregulated in each mutant fish. This will allow for insight about how these differences in gene expression affect the phenotypes observed in this disorder.
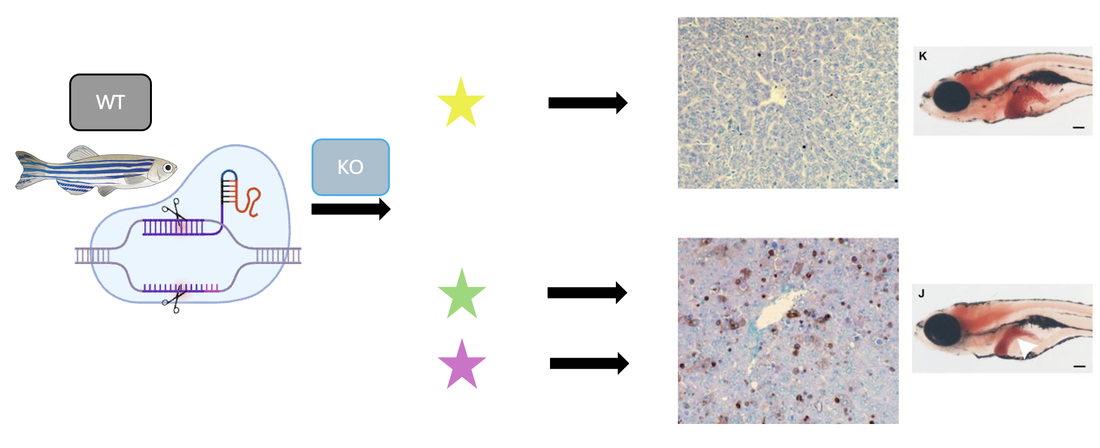
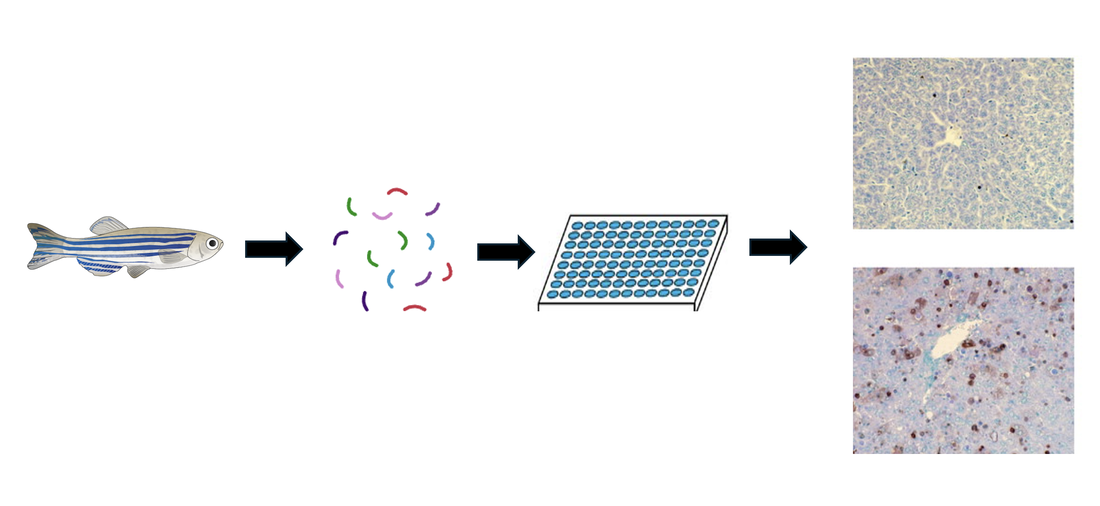
Taking a WT zebrafish and using CRISPR to knockout the three genes observed above, a TUNEL assay was again performed and phenotypes were further analyzed. When the yellow gene was knocked out, the apoptosis levels remained normal and the liver was observed to be healthy and a normal size. When the green and purple genes were knocked out, this TUNEL assay showed a vast increase in apoptosis levels and the decreased liver size can be observed.
From these results, it can be seen that the yellow gene has no impact on the tyrosine catabolism pathway, while the purple and green genes do play a role in the overall regulation of a pathway involved with this disorder. Further research into these genes will allow for a better understanding of the mechanisms involved in healthy cell progression as well as Tyrosinemia Type 1.
aim 3 : quantify differentially expressed proteins in wt and mutant fah hepatocyte cells that lead to apoptosis.
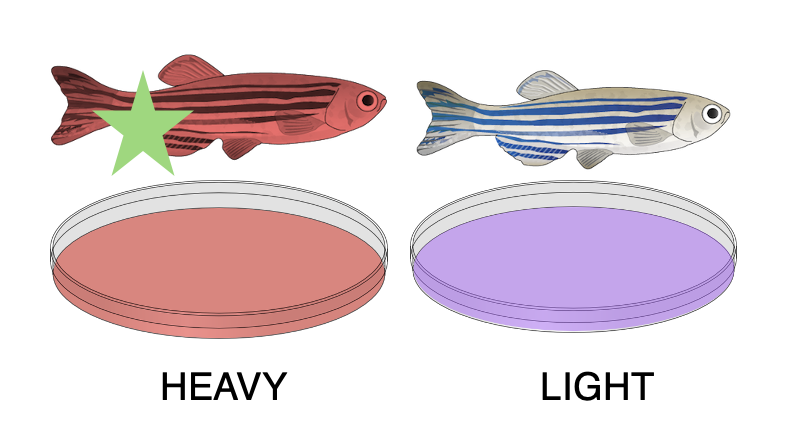
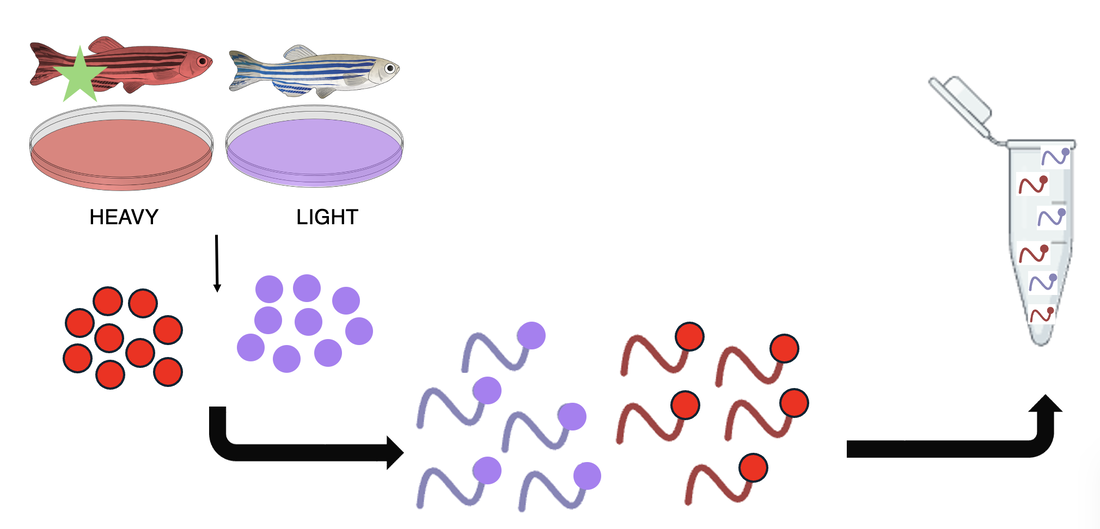
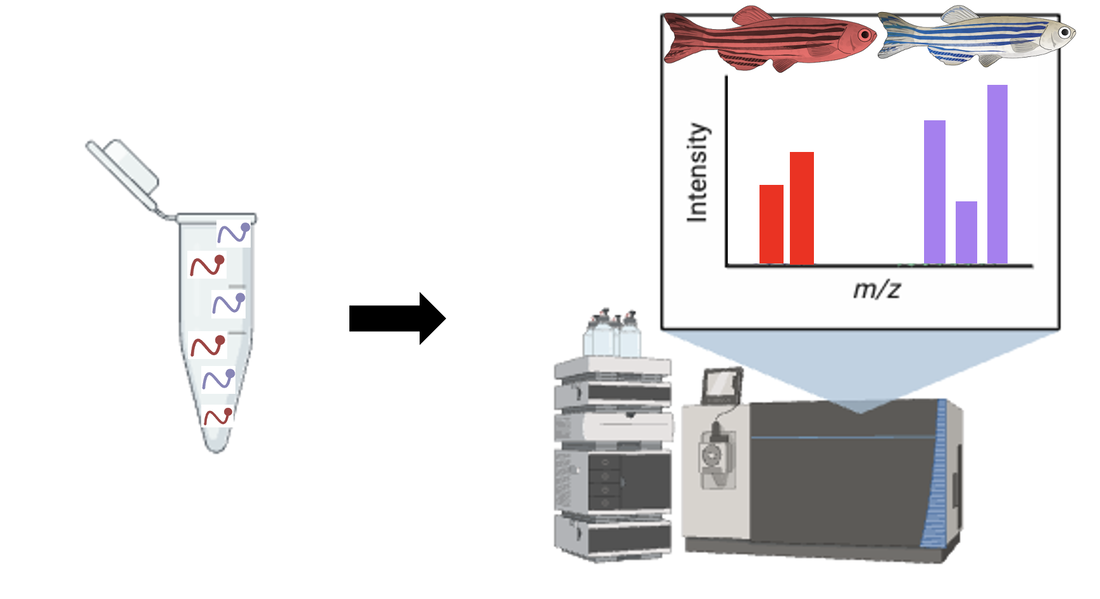
These cells are allowed to grow in the seperate media and are then harvested, lysed, and proteins are extracted. These proteins go through a trypsin process that breaks them into peptides, which can then be mixed together for further analysis.
Putting this mixture through a LC-MS/MS machine, data regarding protein expressions in each fish is produced and separated graphically by the different isotopic weights. Proteins that have different expressions in the mutant fish compared to the WT control group will be used for knockout phenotypic analysis.
Using a WT zebrafish and CRISPR to perform knockouts, proteins that were differentially expressed are knocked out and subjected to a TUNEL assay. The green protein knockout resulted in a normal apoptosis assay and a healthy sized liver. The purple and orange protein knockouts were observed to have an extreme increase in levels of apoptosis and the liver is seen to be much smaller.
This research overall highlighted proteins that are expressed differently when there is a mutation, which is very helpful and can be used in conjugation with a protein interaction network to evaluate different drug treatment options to target these proteins and pathways they are involved in.
conclusions
Overall, these aims will work to highlight different amino acid mutation that are observed to affect liver phenotype and apoptosis levels, as well as differentially expressed genes and proteins between WT and mutant FAH hepatocyte cells. This is extremely useful in understanding how the lack of a functioning FAA enzyme leads to increased apoptosis in hepatocyte cells, and further analysis into these pathways and mechanisms will lead to more drug treatment options in the future.
future research directions
Further chemical genomic research can be done to observe small molecules that interact to create this phenotype. This can be very helpful for seeing drugs that will target this phenotype and stop this increased apoptosis. Research into how a high protein diet affects this disorder would also be very interesting.
final presentation and drafts
| fuerstenau02292024draft1.pdf | |
| File Size: | 3627 kb |
| File Type: | |
| fuerstenau03212024draft2.pdf | |
| File Size: | 10083 kb |
| File Type: | |
| fuerstenau05102024finaltalk.pdf | |
| File Size: | 6929 kb |
| File Type: | |
| fuerstenaufinaltalkanimated.pdf | |
| File Size: | 8170 kb |
| File Type: | |
This web page was produced as an assignment for Genetics 564, a capstone course at UW-Madison.